Variant DetailsVariant: nsv3320410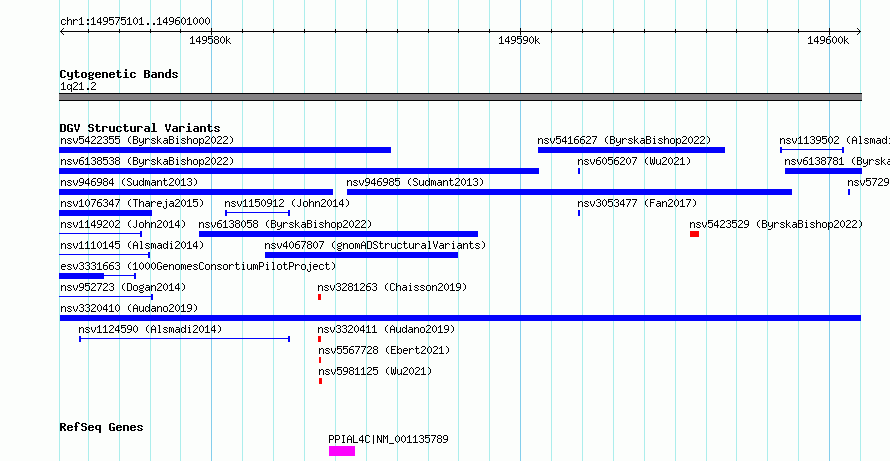 | Internal ID | 19751388 | | Landmark | | | Location Information | | | Cytoband | 1q21.2 | | Allele length | | Assembly | Allele length | | hg38 | 25900 | | hg19 | 772801 |
| | Variant Type | CNV duplication | | Copy Number | | | Allele State | | | Allele Origin | | | Probe Count | | | Validation Flag | | | Merged Status | M | | Merged Variants | | | Supporting Variants | nssv14486220, nssv14475096, nssv14487310, nssv14486647, nssv14484118, nssv14490152, nssv14482237, nssv14491669, nssv14485420, nssv14484980, nssv14491214 | | Samples | HG02106, HG04217, CHM1, NA12878, HG02818, HG02059, HG01352, NA19434, NA19240, HG00733, HG00514 | | Known Genes | FCGR1C, LOC101929780, LOC388692, LOC645166, NBPF23, PPIAL4A, PPIAL4B, PPIAL4C, PPIAL4D, PPIAL4E, PPIAL4F | | Method | Sequencing | | Analysis | Read and contig alignments were done with BLASR. The Celera assembler (8.3rc2) was used to assemble RS II samples (CHM1, CHM13, HG00514, HG00733, NA19240, HG02818, NA19434, HG01352, HG02059, NA12878, and HG04217, and Canu (1.7) was used for Sequel samples (HG02106 and HG00268). Variant calling was performed with SMRT-SV (10.1101/gr.214007.116). | | Platform | PacBio RS II P6C4, PacBio Sequel v2.1 | | Comments | | | Reference | Audano_et_al_2019 | | Pubmed ID | 30661756 | | Accession Number(s) | nsv3320410
| | Frequency | | Sample Size | 14 | | Observed Gain | 11 | | Observed Loss | 0 | | Observed Complex | 0 | | Frequency | n/a |
|
|
