Variant DetailsVariant: nsv1003601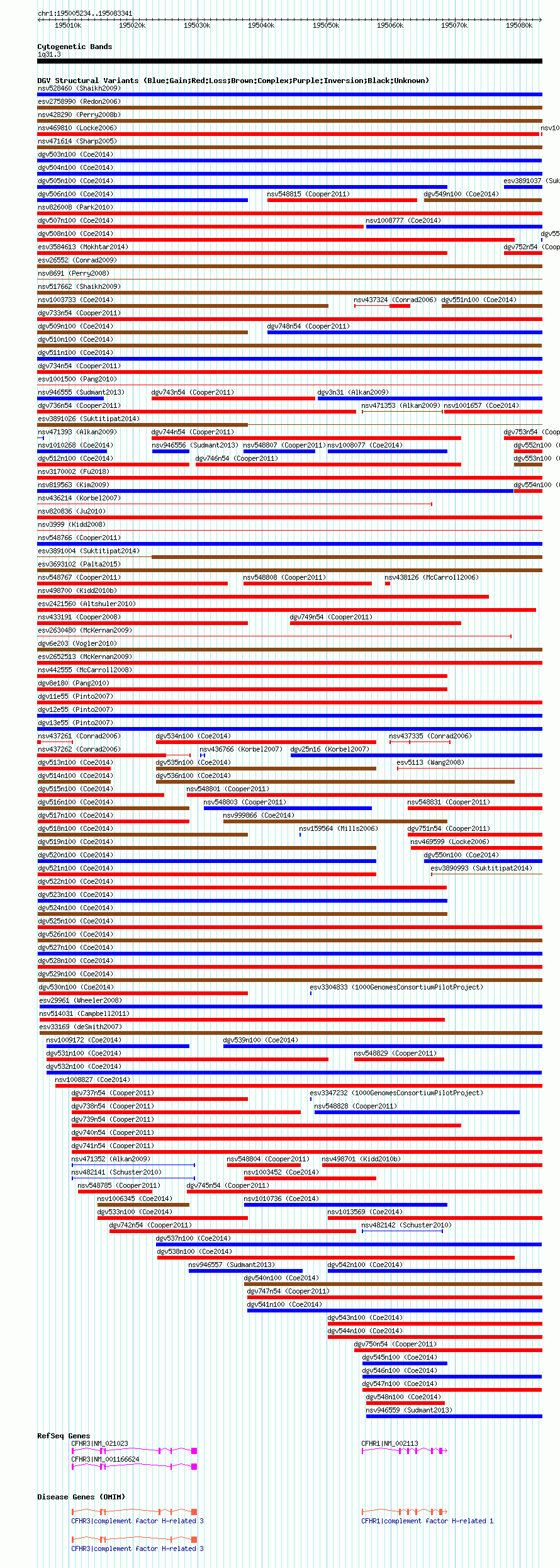 | Internal ID | 19092819 | | Landmark | | | Location Information | | | Cytoband | 1q31.3 | | Allele length | | Assembly | Allele length | | hg38 | 78108 | | hg19 | 78108 | | hg18 | 78108 |
| | Variant Type | CNV gain+loss | | Copy Number | | | Allele State | | | Allele Origin | | | Probe Count | | | Validation Flag | | | Merged Status | M | | Merged Variants | dgv526n100 | | Supporting Variants | nssv3493897, nssv3487787, nssv3489176, nssv3502514, nssv3486486, nssv3494406, nssv3490449, nssv3500229, nssv3487583, nssv3487156, nssv3494668, nssv3487973, nssv3487226, nssv3499329, nssv3498817, nssv3496469, nssv3497575, nssv3497802, nssv3491779, nssv3496623, nssv3486168, nssv3501201, nssv3499625, nssv3498298, nssv3495168, nssv3489696, nssv3493910, nssv3485887, nssv3496161, nssv3500000, nssv3484220, nssv3500652, nssv3496965, nssv3486763, nssv3495977, nssv3485537, nssv3488484, nssv3493204, nssv3488219, nssv3496220, nssv3500131, nssv3483817, nssv3496754, nssv3494765, nssv3486938, nssv3494407, nssv3492886, nssv3490911, nssv3496549, nssv3483995, nssv3484701, nssv3482938, nssv3495537, nssv3705387, nssv3489255, nssv3486485, nssv3488783, nssv3485256, nssv3499124, nssv3493490, nssv3705383, nssv3491132, nssv3489089, nssv3484225, nssv3497099, nssv3490228, nssv3500787, nssv3487603, nssv3502377, nssv3485604, nssv3491878, nssv3499328, nssv3484357, nssv3495320, nssv3494975, nssv3705388, nssv3485836, nssv3502117, nssv3488719, nssv3488487, nssv3486159, nssv3497702, nssv3487137, nssv3498846, nssv3490918, nssv3482920, nssv3483887, nssv3484709, nssv3486311, nssv3490220, nssv3487067, nssv3495484, nssv3485631, nssv3493223, nssv3490422, nssv3705384, nssv3493345, nssv3490832, nssv3485883, nssv3494216, nssv3488586, nssv3492762, nssv3490558, nssv3498563, nssv3484318, nssv3486426, nssv3495024, nssv3493775, nssv3498811, nssv3485920, nssv3483508, nssv3494987, nssv3502373, nssv3495833, nssv3501411, nssv3487189, nssv3495434, nssv3491985, nssv3705389, nssv3496779, nssv3490435, nssv3496876, nssv3497429, nssv3490992, nssv3487097, nssv3492918, nssv3496571, nssv3500934, nssv3705390, nssv3491336, nssv3501950, nssv3488496, nssv3484208, nssv3493435, nssv3490413, nssv3495852, nssv3486284, nssv3493984, nssv3488677, nssv3498366, nssv3486887, nssv3501608, nssv3495786, nssv3498170, nssv3490793, nssv3492825, nssv3486964, nssv3494489, nssv3502374, nssv3490119, nssv3485296, nssv3492130, nssv3705391, nssv3493466, nssv3494960, nssv3490658, nssv3486353, nssv3496891, nssv3705386, nssv3482776, nssv3495798, nssv3492176, nssv3502700, nssv3490700, nssv3494617, nssv3487375, nssv3491952, nssv3493086, nssv3502631, nssv3497245, nssv3492840, nssv3492209, nssv3489587, nssv3485434, nssv3494166, nssv3500407, nssv3495740, nssv3500110, nssv3502089, nssv3497436, nssv3491533, nssv3498481, nssv3487843, nssv3490754, nssv3492290, nssv3500130, nssv3498619, nssv3500903, nssv3490719, nssv3501042, nssv3500244, nssv3497603, nssv3495012, nssv3493560, nssv3499128, nssv3483618, nssv3497585, nssv3491894, nssv3498316, nssv3498271, nssv3499458, nssv3497262, nssv3498090, nssv3496401, nssv3493682, nssv3486262, nssv3491593, nssv3501811, nssv3493011, nssv3490656, nssv3489994, nssv3500756, nssv3488936, nssv3502076, nssv3482818, nssv3494739, nssv3490071, nssv3502623, nssv3497086, nssv3705382, nssv3485926, nssv3494041, nssv3492285, nssv3487591, nssv3494396, nssv3494484, nssv3483123, nssv3488475, nssv3485672, nssv3488444, nssv3491617, nssv3499229, nssv3496734, nssv3705385, nssv3487380, nssv3501788, nssv3493953, nssv3489403, nssv3499209, nssv3493170, nssv3485588, nssv3483389, nssv3494431, nssv3502332 | | Samples | | | Known Genes | CFHR1, CFHR3 | | Method | SNP array | | Analysis | Affymetrix SNP array copy number analysis | | Platform | Affymetrix SNP Array 6.0 | | Comments | | | Reference | Coe_et_al_2014 | | Pubmed ID | 25217958 | | Accession Number(s) | nsv1003601
| | Frequency | | Sample Size | 11257 | | Observed Gain | 177 | | Observed Loss | 67 | | Observed Complex | 0 | | Frequency | n/a |
|
|
