Variant DetailsVariant: nssv544853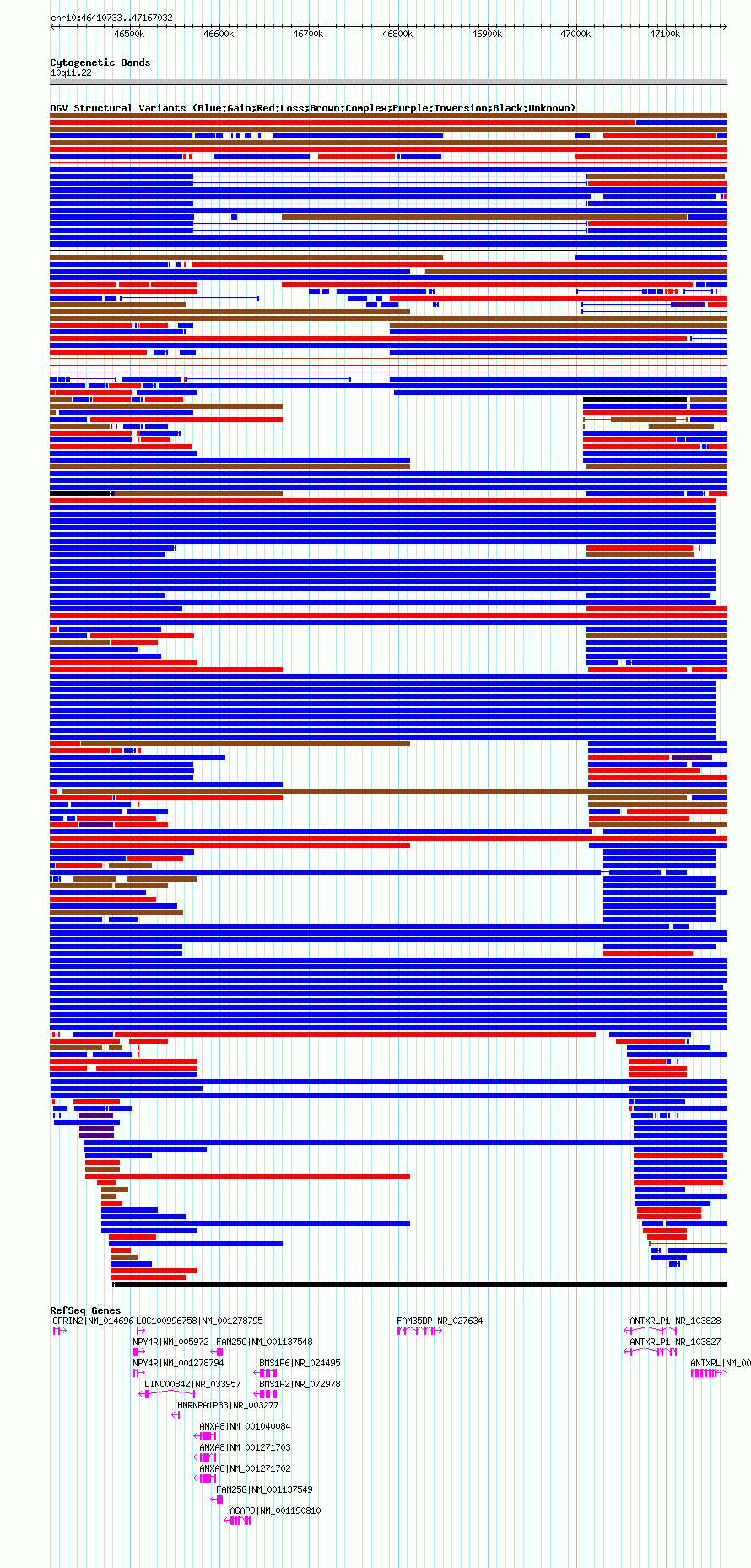 | Internal ID | 15553024 | | Landmark | | | Location Information | | | Cytoband | 10q11.22 | | Allele length | | Assembly | Allele length | | hg19 | 706300 | | hg18 | 756300 |
| | Variant Type | CNV gain | | Copy Number | 4 | | Allele State | | | Allele Origin | | | Probe Count | | | Validation Flag | | | Merged Status | S | | Merged Variants | nsv470934 | | Supporting Variants | | | Samples | HGDP00052 | | Known Genes | AGAP9, ANTXRL, ANTXRLP1, ANXA8, BMS1P2, BMS1P6, FAM25C, FAM25G, FAM35DP, GPRIN2, HNRNPA1P33, LINC00842, LOC100996758, NPY4R | | Method | SNP array | | Analysis | We used the previously validated default quality control criteria, excluding samples with a log R ratio standard deviation of >0.28, a median B allele frequency of >0.55 or <0.45, or a B allele frequency drift of >0.002 (for more details see Wang et al. 2007). As the PennCNV algorithm is more sensitive and specific to CNVs covering greater numbers of SNPs in the HumanHap550 array, use of a minimum number of SNPs in CNV detection increases the reliability of CNV calls (with a consequent reduction in calls per individual). We set 10 SNPs as the minimum detection threshold in the algorithm. | | Platform | Illumina HumanHap550 Genotyping BeadChip v3 | | Comments | Double-copy duplication | | Reference | Jakobsson_et_al_2008 | | Pubmed ID | 18288195 | | Accession Number(s) | nssv544853
| | Frequency | | Sample Size | 443 | | Observed Gain | 1 | | Observed Loss | 0 | | Observed Complex | 0 | | Frequency | n/a |
|
|
