Variant DetailsVariant: esv3578554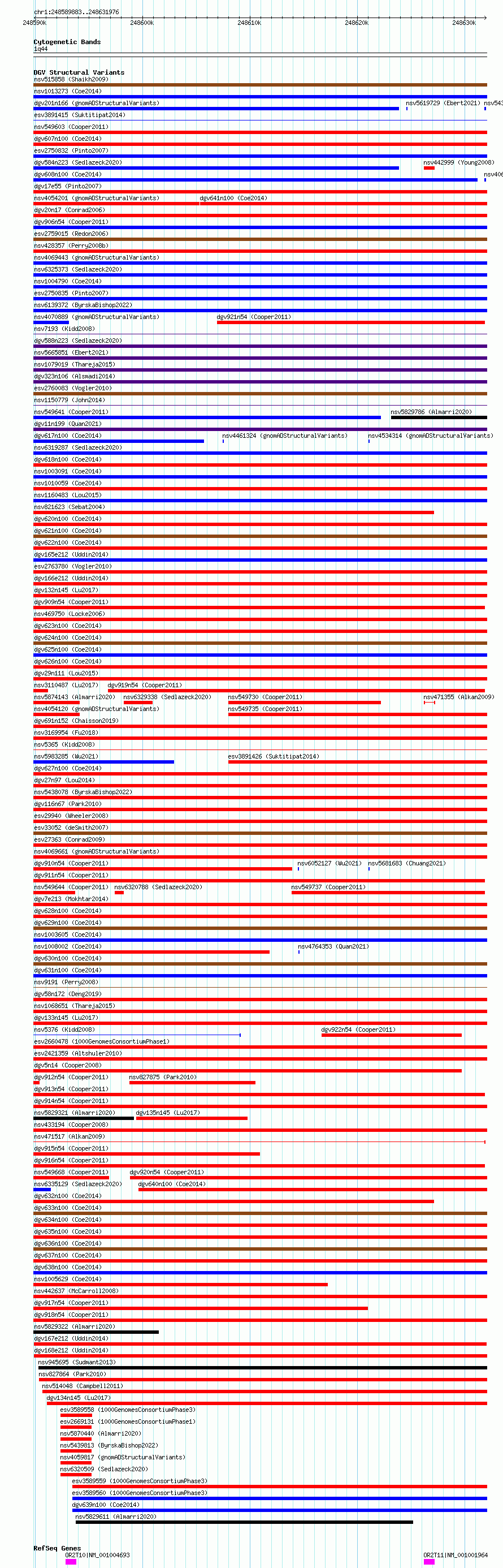 | Internal ID | 18706752 | | Landmark | | | Location Information | | | Cytoband | 1q44 | | Allele length | | Assembly | Allele length | | hg38 | 42094 | | hg19 | 42094 |
| | Variant Type | CNV loss | | Copy Number | | | Allele State | | | Allele Origin | | | Probe Count | | | Validation Flag | | | Merged Status | M | | Merged Variants | dgv167e212 | | Supporting Variants | essv9811744, essv9811500, essv9811234, essv9811645, essv9811578, essv9811378, essv9811733, essv9811323, essv9811556, essv9811411, essv9811767, essv9811478, essv9811678, essv9811278, essv9811700, essv9811711, essv9811778, essv9811367, essv9811212, essv9811589, essv9811511, essv9811633, essv9811545, essv9811334, essv9811534, essv9811756, essv9811345, essv9811423, essv9811811, essv9811312, essv9811434, essv9811600, essv9811300, essv9811256, essv9811356, essv9811722, essv9811622, essv9811656, essv9811489, essv9811389, essv9811400, essv9811800, essv9811245, essv9811467, essv9811223, essv9811522, essv9811267, essv9811289, essv9811567, essv9811611, essv9811689, essv9811789, essv9811456, essv9811201, essv9811445, essv9811667 | | Samples | 400911GA, 401749DJ, 400439IM, 401852SK, 400534ME, 400739SS, 400101EH, 401457WK, 401151RJ, 401949MN, 401093VL, 400855BD, 400509CJ, 401536BD, 401355CD, 400241CP, 401990PR, 400606HW, 400882DD, 400203NA, 402065BG, 400688FL, 400148MS, 401104DM, 401609MB, 401764JJ, 401251WN, 401950MD, 400381CA, 401879HJ, 401630MK, 401075MN, 400888MS, 400639RP, 400248JO, 401795SP, 401514BA, 400518MS, 400837HN, 400103BN, 400053LE, 401277RA, 401894PD, 400586RD, 400072GR, 4000046CJ, 401166WJ, 400792RE, 401543DC, 401105WS, 401177SL, 400508RD, 400300SD, 400238BB, 401490TL, 401246HH | | Known Genes | OR2T10, OR2T11 | | Method | SNP array | | Analysis | We used four separate algorithms to detect CNVs; Affymetrix Chromosome Analysis Suite (ChAS), iPattern, Nexus and Partek. Our primary analysis was performed based on ChAS CNV calls, which were then supported using the remaining three algorithms to construct a confidence set of CNVs. For all algorithms, we have used 8 probes and >1kb as a base line cutoff for CNV detection. | | Platform | Affymetrix CytoScan HD 2.7M array | | Comments | | | Reference | Uddin_et_al_2014 | | Pubmed ID | 25503493 | | Accession Number(s) | esv3578554
| | Frequency | | Sample Size | 873 | | Observed Gain | 0 | | Observed Loss | 56 | | Observed Complex | 0 | | Frequency | n/a |
|
|
