Variant DetailsVariant: esv2662845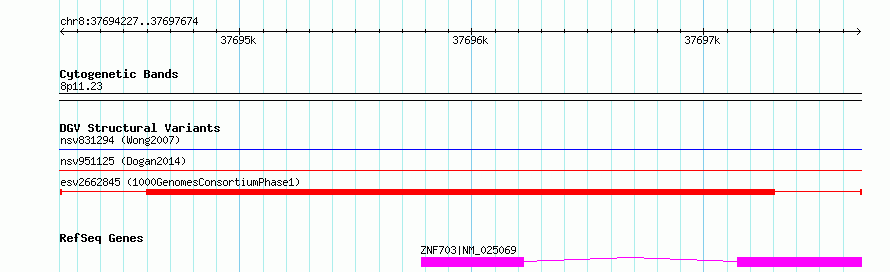 | Internal ID | 9928950 | | Landmark | | | Location Information | | | Cytoband | 8p11.23 | | Allele length | | Assembly | Allele length | | hg38 | 3448 | | hg19 | 3448 |
| | Variant Type | CNV deletion | | Copy Number | | | Allele State | | | Allele Origin | | | Probe Count | | | Validation Flag | | | Merged Status | M | | Merged Variants | | | Supporting Variants | essv5866625, essv6013537, essv5506770, essv6472886, essv6100120, essv5411629, essv5820304, essv6404853, essv5944439, essv5933558, essv6049327, essv5968940, essv5491234, essv6131061, essv5524703, essv5640604, essv6504872, essv6480078, essv6147465, essv5892265, essv5602754, essv5961147, essv6310263, essv6557066, essv6161194, essv5639796, essv5451527, essv5557592, essv6118210, essv6235800, essv5599421, essv6344461, essv5471757, essv6422679, essv5942455, essv6050798, essv5488769, essv6540856, essv5598859, essv6023145, essv5664750, essv5597842, essv5554485, essv5491165, essv5489118, essv6189475, essv6306700, essv6029845 | | Samples | HG00361, HG00318, HG00179, HG00177, HG00327, HG00271, HG00272, HG00330, HG00346, HG00270, HG00185, HG00281, HG00277, HG00335, HG00325, HG00309, HG00182, HG00326, HG00178, HG00323, HG00313, HG00268, HG00266, HG00176, HG00282, HG00328, HG00368, HG00320, HG00344, HG00284, HG00273, HG00373, HG00276, HG00336, HG00285, HG00353, HG00357, HG00319, HG00339, HG00269, HG00312, HG00329, HG00342, HG00174, HG00280, HG00343, HG00274, HG00171 | | Known Genes | ZNF703 | | Method | Merging | | Analysis | No reference, merging analysis | | Platform | Merging | | Comments | High quality site | | Reference | 1000_Genomes_Consortium_Phase_1 | | Pubmed ID | 23128226 | | Accession Number(s) | esv2662845
| | Frequency | | Sample Size | 1151 | | Observed Gain | 0 | | Observed Loss | 48 | | Observed Complex | 0 | | Frequency | n/a |
|
|
