Variant DetailsVariant: essv9793343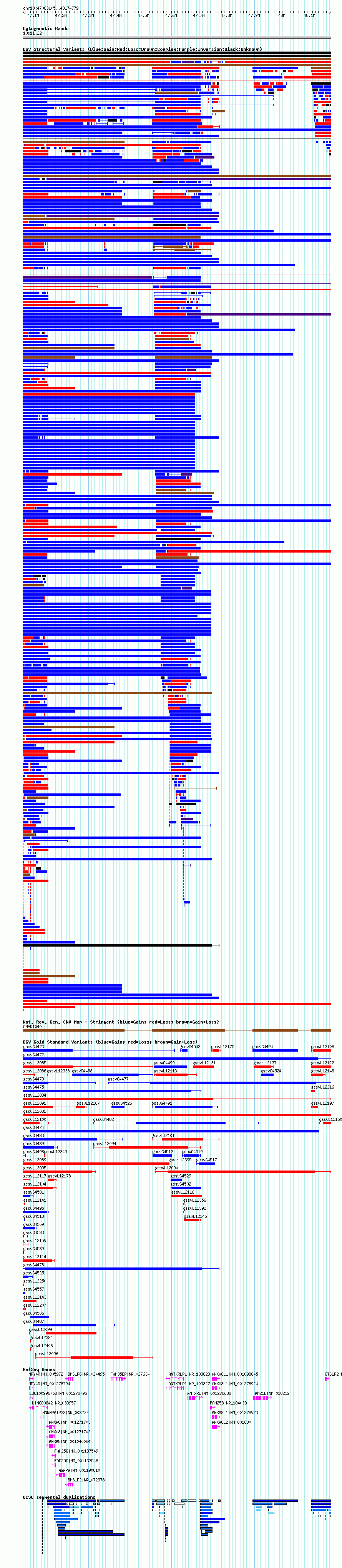 | Internal ID | 18629510 | | Landmark | | | Location Information | | | Cytoband | 10q11.22 | | Allele length | | Assembly | Allele length | | hg19 | 1111675 |
| | Variant Type | CNV loss | | Copy Number | 1 | | Allele State | | | Allele Origin | | | Probe Count | | | Validation Flag | | | Merged Status | S | | Merged Variants | esv3578846 | | Supporting Variants | | | Samples | 400114GR | | Known Genes | AGAP9, ANTXRL, ANTXRLP1, ANXA8, ANXA8L1, ANXA8L2, BMS1P2, BMS1P6, CTSLP2, FAM21B, FAM25B, FAM25C, FAM25G, FAM35DP, HNRNPA1P33, LINC00842, LOC100996758, NPY4R | | Method | SNP array | | Analysis | We used four separate algorithms to detect CNVs; Affymetrix Chromosome Analysis Suite (ChAS), iPattern, Nexus and Partek. Our primary analysis was performed based on ChAS CNV calls, which were then supported using the remaining three algorithms to construct a confidence set of CNVs. For all algorithms, we have used 8 probes and >1kb as a base line cutoff for CNV detection. | | Platform | Affymetrix CytoScan HD 2.7M array | | Comments | Number of probes=147 | | Reference | Uddin_et_al_2014 | | Pubmed ID | 25503493 | | Accession Number(s) | essv9793343
| | Frequency | | Sample Size | 873 | | Observed Gain | 0 | | Observed Loss | 1 | | Observed Complex | 0 | | Frequency | n/a |
|
|
