Variant DetailsVariant: essv18489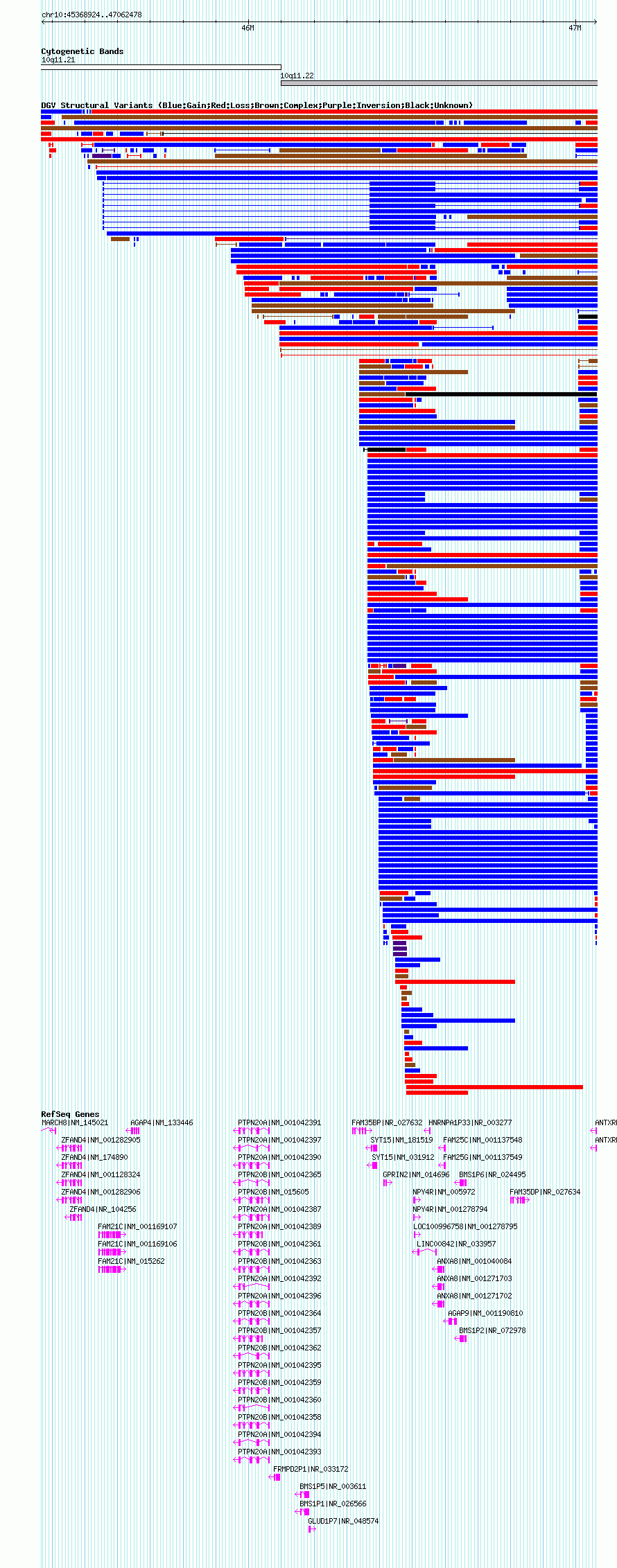 | Internal ID | 9958541 | | Landmark | | | Location Information | | | Cytoband | 10q11.21 | | Allele length | | Assembly | Allele length | | hg38 | 156348 | | hg19 | 1543555 | | hg18 | 1693555 | | hg17 | 1693555 |
| | Variant Type | CNV gain | | Copy Number | | | Allele State | | | Allele Origin | | | Probe Count | | | Validation Flag | | | Merged Status | S | | Merged Variants | esv2757382 | | Supporting Variants | | | Samples | NA12156 | | Known Genes | AGAP4, AGAP9, ANTXRLP1, ANXA8, BMS1P1, BMS1P2, BMS1P5, BMS1P6, FAM21C, FAM25C, FAM25G, FAM35BP, FAM35DP, FRMPD2P1, GLUD1P7, GPRIN2, HNRNPA1P33, LINC00842, LOC100996758, MARCH8, NPY4R, PTPN20A, PTPN20B, SYT15, ZFAND4 | | Method | SNP array | | Analysis | The algorithm used to call CNVs using the 500K EA platform was developed to accurately define CNV regions using a large set of reference samples and is described in detail in a separate publication (Komura 2006). The algorithm contains three major parts: 1) Intensity pre-processing using an improved version of Genomic Imbalance Map (GIM) (Ishikawa et al. 2005), including probe selection, noise reduction, normalization, and intensity ratio adjustment based on affinity differences between alleles of a SNP, 2) CNV extraction, which identifies CNVs from all pair-wise comparisons using a modified SW-ARRAY, and 3) A copy number inference step which utilizes signal ratios and SNP information to more precisely define CNV boundaries and the copy number within each region. | | Platform | Affymetrix GeneChip Early Access Mapping 500K Set Array (250K_Nsp_SNP) | | Comments | | | Reference | Redon_et_al_2006 | | Pubmed ID | 17122850 | | Accession Number(s) | essv18489
| | Frequency | | Sample Size | 270 | | Observed Gain | 1 | | Observed Loss | 0 | | Observed Complex | 0 | | Frequency | n/a |
|
|
