Variant DetailsVariant: dgv66n17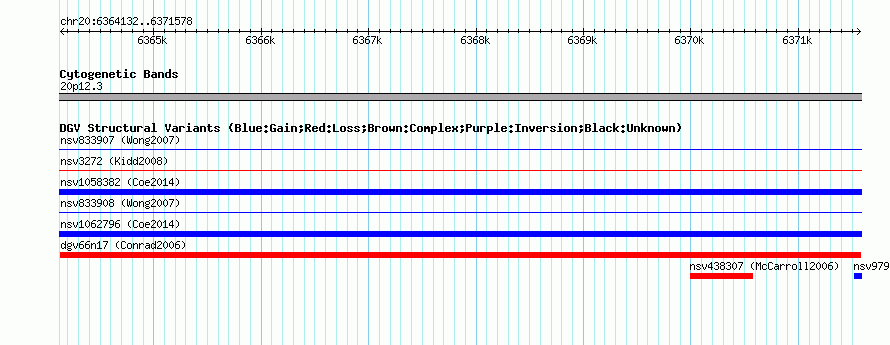 | Internal ID | 22766103 | | Landmark | | | Location Information | | | Cytoband | 20p12.3 | | Allele length | | Assembly | Allele length | | hg38 | 7447 | | hg19 | 7447 | | hg18 | 7447 | | hg16 | 7447 |
| | Variant Type | CNV loss | | Copy Number | | | Allele State | | | Allele Origin | | | Probe Count | | | Validation Flag | | | Merged Status | M | | Merged Variants | | | Supporting Variants | nsv437837, nsv437836 | | Samples | NA19194, NA18863 | | Known Genes | | | Method | SNP array | | Analysis | Our algorithm aims to detect deletions that are transmitted from a hemizygous parent to a child. For each trio, every SNP was coded into one of seven categories: (A) Type I mendelian incompatibility (that is, consistent with deletion) involving mother; (B) Type I mendelian incompatibility involving father; (C) Type II mendelian incompatibility (that is, inconsistent with deletion); (D) child homozygous or missing data, both parents homozygous or missing data; (E) child homozygous or missing data, father heterozygous, mother homozygous or missing data; (F) child homozygous or missing data, mother heterozygous, father homozygous or missing data; (G) child heterozygous or both parents heterozygous (see Supplementary Methods for further details). SNPs were assigned to states D-G only if they did not contain mendelian incompatibilities. A run of consecutive SNPs in a particular trio was considered to be consistent with a maternal transmitted deletion if all SNPs were in states A, D or E, or with a paternal deletion if all SNPs were in states B, D or F. | | Platform | Not reported | | Comments | | | Reference | Conrad_et_al_2006 | | Pubmed ID | 16327808 | | Accession Number(s) | dgv66n17
| | Frequency | | Sample Size | 60 | | Observed Gain | 0 | | Observed Loss | 2 | | Observed Complex | 0 | | Frequency | n/a |
|
|
